| C-Tyr
Cytosine-1-yl-(2-carboxyethyl)-L-tyrosine 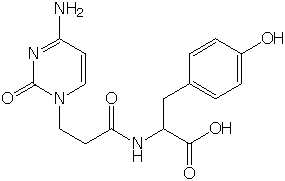
Crystal Structure of Hybrid Dipeptide, Cytosinyl-L-Tyrosine Mitsunobu DOI, Hajime MIYAKO, Akiko ASANO and Toshimasa ISHIDA Analytical Sci., (1999) 15, 109-110. This study was supported by the Grant-in-Aid for Scientific Research 07672427 from the Ministry of Education, Science, Sports and Culture. |
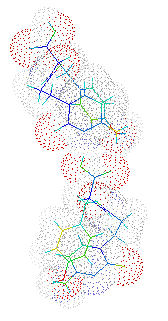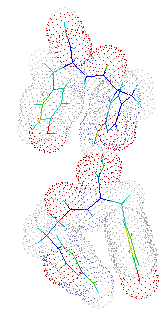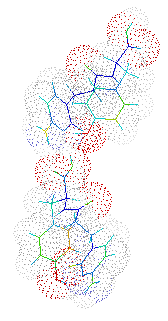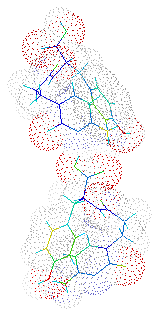 This structer was produced by RasMol, and stucked by GifBuilder (Yves Piguet, 1997) |
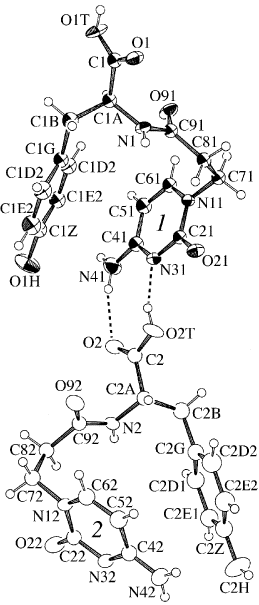 ORTEP-III (Burnett, 1996) drawing |
Hybrid dipeptide was designed to investigate the cooperative interactions
between the nucleic acid and polypeptide,1 and the structure property is
similar to that of peptide nucleic acid (PNA). In this paper, we report
the interaction between tyrosine and cytosine. The title compound
(C-Tyr) was synthesized by the coupling 1-ethylcarboxy-cytosine and
tyrosine methyl ester, and crystals were obtained from an aqueous methanol
solution. There are two crystallographically independent molecules
(Molecules 1 and 2) in the asymmetric unit and are discriminated by the
differences in the torsion angles around C8-C9, N-CA and CB-CG which
are related with the oriantations of aromatic rings. The molecules are
folded with the similar mannerand, and the overall conformations are
similar to each other. The turned conformation results the access between
the cytosine base and the phenol ring, though no intramolecular
interaction is observed. The C-terminal groups of tyrosine seem to be an
unionized form (-COOH) in both molecules since the bond distances of C-O
and C-OT are significantly differed. The cytosine bases interacte with
these carboxyl groups: N41...O2, O2T...N31, N42...O1 and O2T...N31. In
addition to these hydrogen bonds, the C-H...O hydrogen bonds are observed.
In the C-Tyr crystal, the distances of H5...O9 and H6...O9 are below the
sum of the van der Waals radii of hydrogen and oxygen (=2.7 A).
|
X-Ray Data Summary
| formula | 2(C16 H18 N4 O5) | ||
| weight | 692.684 | ||
| symmetry | monoclinic | ||
| space group | P21 | ||
| Cell | Crystal | ||
| a (Ang) | 9.5626(12) | description | plate |
| b (Ang) | 10.4539(12) | colour | colorless |
| c (Ang) | 16.864(6) | size (mm) | 0.70x0.12x0.05 |
| alpha (deg) | 90.0 | Dx (g/ml) | 1.367 |
| beta (deg) | 93.218(17) | F(000) | 728 |
| gamma (deg) | 90.0 | mu(CuKa) | 0.870 |
| V (Ang^3) | 1683.1(6) | ||
| Z | 2 | ||
| Refinement | Diffrn measurement | ||
| number_reflns | 2841 | device type | Rigaku AFC5R/RU-200 |
| parameters | 291 | decay | 3.0 % |
| restraints | 0 | index limit | 0-h-11,-12-k-0,-19-L-19 |
| R_factor_all | 0.0703 | theta (deg.) | 2.62 - 64.87 |
| R_factor_gt | 0.0661 | total reflections | 2841 |
| wR_factor_ref | 0.1890 | reflections(obs) | 2721 .gt. 2sigma(I) |
| wR_factor_gt | 0.1849 | Structure | |
| shift/su_max | lt.0.001 | solution | SHELXS-97 |
| shift/su_mean | lt.0.001 | refinement | SHELXL-97 |
Get PDB coordinates
Get CIF text
Get SHELXL INS file
 Back to the structure index
Back to the structure index
Date: Jly.1998